This is a 2002 webpage of Pellionisz, when his (now issued) 8,280,641 patent application was filed. (In 2002 heralded here)
This webpages is linked from the book by Perry Marshall (2015), "EVOLUTION 2.0" (page 281).
The impact on theories of evolution was obvious at the "heureka! moment of FractoGene ("Fractal Genome Grows Fractal Organisms").
Cerebellar Purkinje brain cells are fractal (in black, on the right side of diagram below).
Both the evolution of the complexity of Purkinje brain cells, and its correlation with the non-coding DNA repeats was both predicted by FractoGene,
and was experimentally proven in a quantitative and visual manner (Simons and Pellionisz, 2006).
Perry Marshall destroys the misconception of "randomness", and substitutes it by "fractal" on page 281 of his book.
Perry Marshall cites in his ample set of references for the "mathematics" by Pellionisz et al. (2013).
For an overview of FractoGene Decade see a set of links, and Professional Website of Andras Pellionisz.
[2002]
*** ARE YOU 99% MOUSE? THE EVOLUTION REVOLUTION ***
AS AN EMBRYO YES, YOU ARE. IN YOUR GENES, YES, YOU ARE.
IN YOUR INTRONS, NO, YOUR ARE NOT.
INTRONS HELP MAKING US HUMAN, BUT ... PLEASE EXPLAIN HOW?
Aspects of evolution of FractoGene utility patent are provided for illustrative purposes. From a patent the requirements are that it must "theach new skills". FractoGene by measuring and comparing non-coding sequences of DNA as fractal sets, does that. However, from the enormous publicity that FractoGene generated it is evident that there is a pressure from Academia to learn as much about the science background of the patent as possible - without compromising the business interests. The author, having done his "homework" in Academia, is open to all classic academic interests, presentations, publications, etc. This page is a preview to show that FractoGene is not only consistent with prevailing scientific facts of evolution, but actually explains more, and the explanation is clearer (quantitative) - as it is mathematical. Therefore, quantitative predictions will both stimulate experimentation as well as facilitate the profitability of a utility patent.
Thomas Edison used to say for the accomplishments of an inventor: "It takes 1% inspiration, 99% perspiration"
Given this classic slogan, we should not be overly surprised, that up to 97% of the human genetic information (DNA) is seemingly needless, repetitive "junk" - only about 3% is known to generate proteins, deserving the name "gene". The rest used to be called "junk DNA", lately renamed as "non-coding DNA", sometimes labeled by the mysterious, though not very explicit description that these self-similar strands "regulate gene expression".
This uncertainity if "junk DNA" is really "junk", that cought some scientists by surprise, is now history. Indeed, scientists were never quite sure that any part of DNA could be discarded outright, and even the question was raised if introns might play a role in evolution, see Mattick, 1994., Klyce, 2002). The bombshell of the 5th of December issue of the highly reputed Nature 420, 509 (2002) revealed that the 99% of the genes of the mouse have direct equivalents with the humane genes.
What does this mean?
1) It does not mean that homo sapiens is 99% identical to mouse. It has also been revealed, that the mouse DNA is at least 10% less information, compared to the human DNA.
2) The significant difference is less visible in the "inspiration" to make a human out of a lower mammal. It is clearly the "perspiration" that dominates the effort. It is the introns.
3) Are there any clear, maybe even mathematical explanations available how the repeated "non-coding sequences" refine the basic "invention" expressed by a gene?
FractoGene, the patent application authored and submitted by July 31st, 2002, does provide the following explanation:
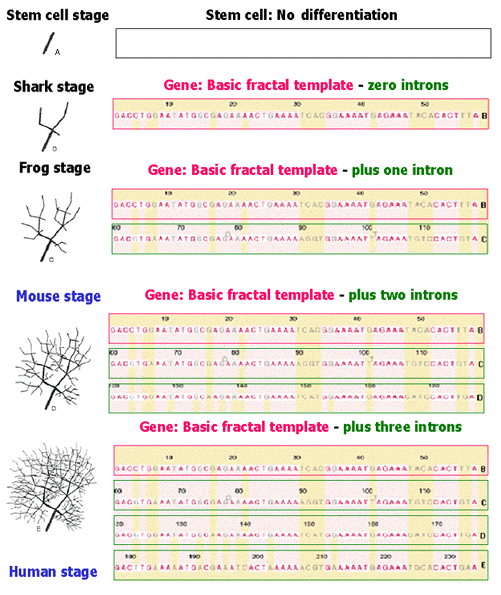
FractoGene patent (pending) pictures evolution as fractal elaboration of a gene
by its increasing amount of non-coding DNA (the fractal model of Purkinje cell was published by Pellionisz, 1989)
The stem (as in "stem cell") in Fig. A. shows no differentiation. In Fig. B., the basic row of base-pair sequence, composed of an ACTG jumble (a "gene") determines the fractal template of the bifurcation at the rudimentary, let's call it "shark" stage. (The cerebellum is generally regarded to be the "invention" of the sharks about 400 million years ago - but the evolutionary "root" has yet to be conclusively established, and as shark DNA has not been sequenced, thus it is unknown if the "gene" determining the shape of Purkinje cells, contains much fewer or no introns, compared to higher species). The repetition of the gene in a self-similar intron (which is not identical to the gene, since the branchlets of the generated brain cell are replaced by the template in e.g. a smaller version as shown in C). For the sake of simplicity, (given the compressed scheme of the diagram), let's call it "Frog stage". Suppose, that Mother Nature is content with such a primitive rendering, e.g. for the frog. In this case the lower-ranking animal only needs two self-similar strands of base-pair sequences (one gene and one intron). Let's assume for the sake of a little more complexity, that evolution does not stop at the "frog stage". A third row of base-pair sequence replaces each terminal branchlet of C with the fractal template itself (in an even smaller version, plus e.g. with a higher density of synapses). This could result in the Purkinje cell of a creature somewhat higher on the evolutionary ladder from the "frog", let us call it symbolically "a mouse". For this to happen, a third repetition, one gene and two introns, "non-coding sequences" are needed, generating D. With yet another round of "perspiration", the outer branchlets of D are replaced by the tiny version of the basic fractal template shown in A, and thus E arises. For the sake of this diagram, let's figuratively call it "Human stage" (although a Purkinje cell at this level of complexity approximates only the guinea pig). As shown the bottom Purkinje cell (and in the original publication), this is a strinkingly accurate fractal model of a fairly sophisticated Purkinje neuron (E).
Why is FractoGene called here to help an "evolution revolution"? Because a paradigm-shift is happening, a transition from the classic notion that the main drivers of evolution are the "genes". Apparently, the primary drivers are the repetitive, "non-coding" introns, constantly improving on the "performance" of the basic "inventions" of the genes. In view of this schematic diagram of fractogenesis (evolution expressed in fractal geometry), it is evident that the "gene pool" of the mouse and human is 99% identical (as found by the Consortium, and reported by Nature, Dec. 5th, 2002) - while there is a fairly significant difference of the total volume of the DNA, it is an experimentally established trend that the higher ranking species contain significantly more "introns".
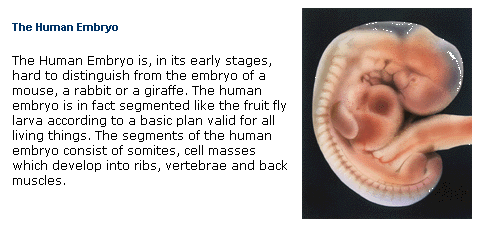
From the schematic diagram based on the FractoGene patent, it is also evident why the development of the embryo "mimics" the development of species through the evolution, both embryogenesis repeating phylogenetic development in general, (see excerpt Figure below), and research is intensifying to explore this phenomenon at genomic and also at the molecular levels.